How Human Cells Read a Hidden Layer of the Genetic Code
Kyoto University / Science
The genetic code has a curious feature that has puzzled molecular biologists for decades. Multiple three-letter sequences, called codons, can encode the same amino acid. On paper, these synonymous codons should be interchangeable. They are not.
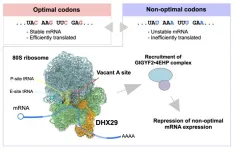
Some codons are translated efficiently by the cell's protein-making machinery. Others are sluggish, causing the ribosome to stall. Messenger RNAs loaded with these "non-optimal" codons tend to be unstable and get degraded. But how human cells actually detect the difference between a fast codon and a slow one has remained largely unknown. Until now.
Finding the sensor
A collaborative team from Kyoto University and RIKEN, led by Osamu Takeuchi and Takuhiro Ito, set out to identify the molecular machinery responsible for reading this hidden layer of information in the genetic code. Their approach was systematic and multi-pronged.
First, they ran a genome-wide CRISPR screen, knocking out genes one by one to see which ones affected codon-dependent gene expression. The screen pointed them to a single RNA-binding protein: DHX29.
When the team deleted DHX29 from cells and used RNA sequencing to measure global mRNA levels, they found something telling. Without DHX29, mRNAs enriched in non-optimal codons were no longer being degraded as efficiently. They accumulated. DHX29, it turned out, was not just involved in codon quality control. It was central to it.
Watching the protein at work on the ribosome
To understand how DHX29 operates mechanically, the researchers turned to cryo-electron microscopy, a technique that freezes molecular complexes in place and images them at near-atomic resolution. The resulting structures showed DHX29 sitting directly on the 80S ribosome, the cell's protein-synthesis machine.
Selective ribosome profiling then revealed a key detail about DHX29's preferences. The protein did not bind ribosomes randomly. It preferentially associated with ribosomes that were stalled on non-optimal codons, exactly where you would expect a quality-control sensor to position itself.
The final piece of the puzzle came from proteomic analyses, which showed that DHX29 recruits a partner complex called GIGYF2-4EHP. This complex acts as a brake, selectively repressing translation of mRNAs that contain too many non-optimal codons. The chain of events runs from codon recognition to ribosome binding to translational repression and mRNA degradation. It is a complete surveillance pathway.
Why codon choice matters beyond the textbook
The implications stretch well beyond basic molecular biology. Codon usage varies across different genes and different cell types. During cell differentiation, when a stem cell commits to becoming a neuron or a muscle cell, the pool of available transfer RNAs shifts. What counts as an "optimal" codon in one cell type may be non-optimal in another. DHX29's quality-control role could therefore influence which proteins are made, and in what quantities, as cells change their identity.
There are also potential links to disease. Cancer cells are known to alter their codon usage patterns, and disruptions to mRNA quality control have been implicated in tumor development. If DHX29 plays a role in suppressing the expression of genes with non-optimal codons, losing that function could allow aberrant proteins to accumulate.
Co-corresponding author Masanori Yoshinaga noted that these findings establish a direct molecular link between synonymous codon choice and gene expression control in human cells, a connection that was theorized but lacked a defined mechanism.
What remains uncertain
The study was conducted primarily in human cell lines, and the team has not yet demonstrated how DHX29 loss affects a living organism. Whether the protein plays equally important roles across all tissue types, or whether its influence is concentrated in rapidly dividing cells, remains to be tested. The connection to cancer is plausible but unproven; it will require disease models to move from hypothesis to evidence.
The team's next step is to explore how DHX29's influence on gene expression translates to health and disease outcomes. Takeuchi described the discovery as particularly satisfying after years of investigating how cells interpret the information embedded within the genetic code.