Bacteria separate their DNA using accordion-like folds and molecular repulsion
Rice University. Supported by NSF (PHY-2019745, PHY-2210291, PHY-2014141) and Welch Foundation (C-1792).
Bacterial chromosomes fold themselves like an accordion during replication, and that folding generates enough repulsive force to shove two copies of the genome to opposite sides of the cell. That's the central finding from a new study by physicists at Rice University, and it answers a question that has nagged biophysicists for years: how do bacteria separate their DNA without the elaborate spindle apparatus that human cells build for the job?
No spindles, no problem
Human cells divide through mitosis, a carefully choreographed process in which protein spindles physically grab chromosomes and pull them apart. Bacteria skip all of that. They reproduce through binary fission — a faster, more streamlined process in which the circular chromosome begins separating even as it's being copied. One cell becomes two, each with its own complete genome, in a fraction of the time mitosis requires.
But speed and simplicity raise a mechanical question. If nothing is pulling the chromosomes apart, what pushes them? Jose Onuchic, the Harry C. and Olga K. Wiess Chair of Physics at Rice and the study's corresponding author, wanted to understand the forces at work.
Building a chromosome model from Hi-C data
The research team turned to Hi-C maps — experimental datasets that reveal how chromosomes are organized in three-dimensional space by measuring which regions of DNA are physically close to each other. By combining Hi-C data with physical modeling, first author Sumitabha Brahmachari and colleagues constructed detailed three-dimensional models of bacterial chromosomes at each stage of replication.
The critical comparison came from running the model twice: once with fully functional SMC proteins (structural maintenance of chromosomes, a highly conserved protein family found across bacterial species) and once with a defective version. Watching how chromosome architecture evolved over time, with and without SMC, revealed what these proteins actually do during replication.
The accordion mechanism
Bacterial DNA replication starts at a specific point on the circular chromosome called the origin of replication, or ori. From there, the copying machinery moves in both directions simultaneously around the circle. Picture a clock with the ori at twelve o'clock — replication proceeds clockwise and counterclockwise at the same time, eventually meeting at six o'clock.
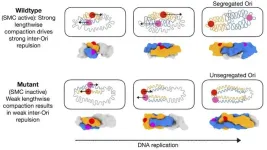
When SMC proteins are present, the newly copied DNA doesn't just trail loosely behind the replication machinery. It gets compacted lengthwise — folded into tight, accordion-like pleats. This compaction is the key. The folded DNA occupies less linear space but generates repulsive forces between the two copies. The more DNA gets replicated and compacted, the stronger the repulsion becomes, and the further apart the two ori regions drift.
By the time replication reaches the halfway point, the model shows enough accumulated repulsion that the new copy begins peeling away from the original. When replication completes, the two ori regions sit on opposite sides of the cell. The cell membrane pinches inward between them, dividing neatly into two daughter cells, each with one chromosome.
Without SMC, the copies collapse
The model without functional SMC proteins tells a starkly different story. Repulsive forces between the two DNA copies still exist — they're an inherent consequence of having two large polymers in a confined space — but they're far weaker. Instead of accordion folds generating robust separation, the DNA copies collapse into flexible, tangled states. The ori regions stay close together rather than migrating to opposite poles.
The consequences for the cell are severe. Without proper separation, binary fission can damage the DNA as the cell tries to divide. One daughter cell might end up with two copies of the chromosome while the other gets none. The process that should be reliable and fast becomes error-prone.
Forces, not structures
What makes this finding notable is that bacteria achieve faithful chromosome segregation through emergent physical forces rather than dedicated molecular machinery. Human cells build an entire apparatus — the mitotic spindle, with its kinetochores and motor proteins — to pull chromosomes apart. Bacteria generate separation as a byproduct of how SMC proteins organize DNA during replication. The folding creates the force. The force creates the separation. No external structure required.
The study doesn't resolve how SMC proteins produce the accordion-like compaction in the first place. That mechanism — the molecular details of how these ring-shaped proteins thread and fold DNA — remains an active area of investigation. The tangled states that emerge in SMC-deficient bacteria also need further characterization.
Colony speed and the physics underneath
Bacteria are under evolutionary pressure to replicate quickly. Colony growth depends on it. Binary fission, at its best, is a process optimized for speed: simultaneous replication and segregation, no waiting for a spindle to assemble, no checkpoint delays. But that speed depends on the physics working correctly — on SMC-driven compaction generating enough repulsive force to separate chromosomes reliably every time.
The Rice team's model provides the framework for understanding those forces quantitatively. Future work can now use this framework to probe additional questions: how does the process adapt when bacteria are stressed or nutrient-deprived? What happens during overlapping rounds of replication, when fast-growing bacteria begin copying their DNA before the previous round finishes? How do the forces scale across different bacterial species with different chromosome sizes?
The study was limited to computational modeling validated against Hi-C data, not direct experimental visualization of the separation process in live cells. The models describe what happens and predict the forces involved, but the precise molecular choreography of SMC on DNA at the single-molecule level remains to be captured.